Transcription shapes DNA replication initiation and termination in human cells. Dual roles of poly(dA:dT) tracts in replication initiation and fork collapse. Pif1-family helicases cooperatively suppress widespread replication-fork arrest at tRNA genes. Replication landscape of the human genome. 
Genome-wide nucleotide-resolution mapping of DNA replication patterns, single-strand breaks, and lesions by GLOE-Seq. Spatiotemporal coupling and decoupling of gene transcription with DNA replication origins during embryogenesis in C. Chromosomal ARS1 has a single leading strand start site. Intrinsic coupling of lagging-strand synthesis to chromatin assembly. Detection and sequencing of Okazaki fragments in S. Possible discontinuity and unusual secondary structure of newly synthesized chains. Okazaki, R., Okazaki, T., Sakabe, K., Sugimoto, K. Peaks cloaked in the mist: The landscape of mammalian replication origins. Stalled fork rescue via dormant replication origins in unchallenged S phase promotes proper chromosome segregation and tumor suppression. Dormant origins licensed by excess Mcm2-7 are required for human cells to survive replicative stress. 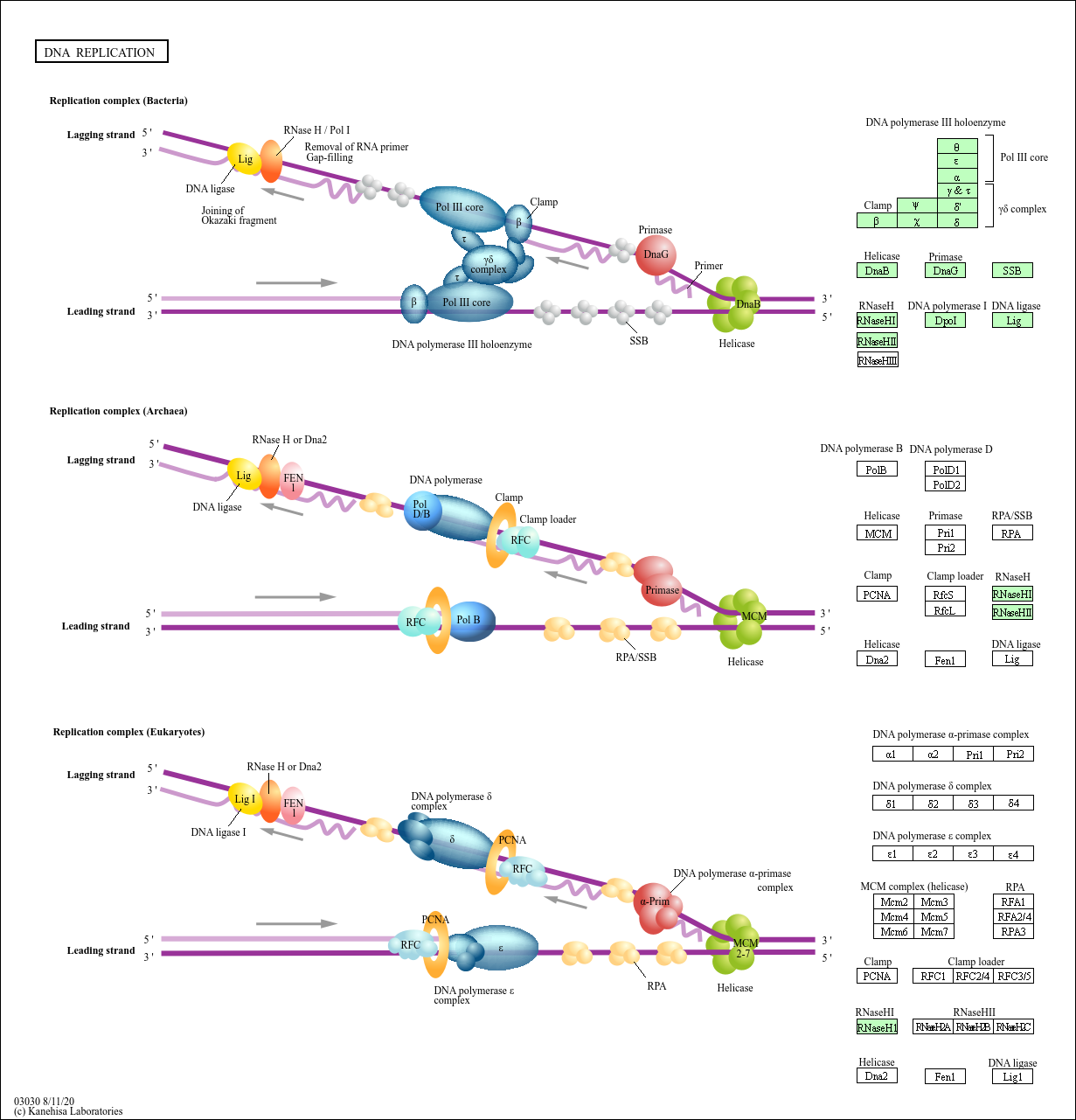
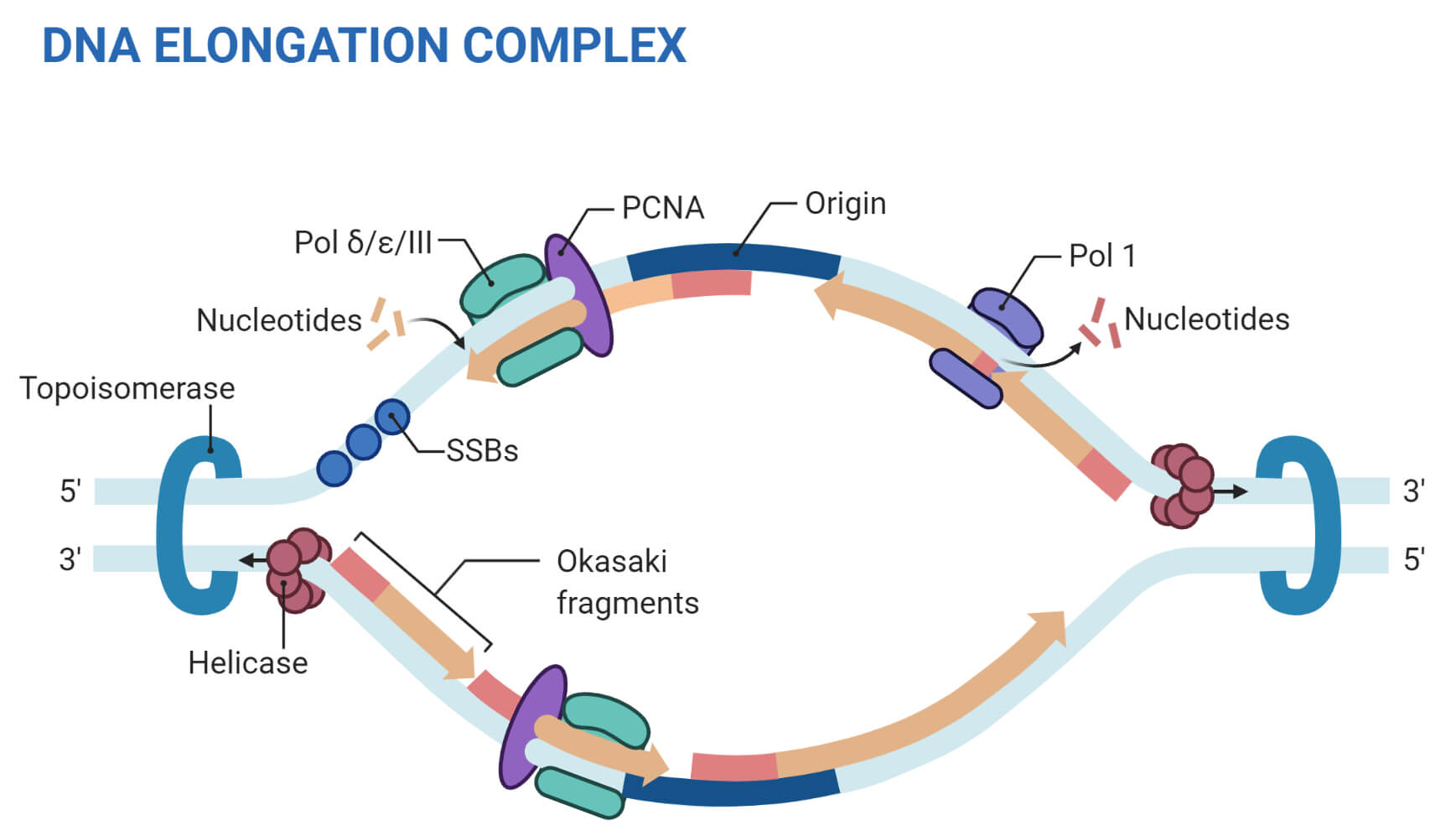
Dormant origins, the licensing checkpoint, and the response to replicative stresses. Excess MCM proteins protect human cells from replicative stress by licensing backup origins of replication. DNA replication origin activation in space and time. DNA replication stress as a hallmark of cancer. The use of Ok-seq to interrogate genome-wide replication fork initiation and termination efficiencies can be applied to all unperturbed, asynchronously growing mammalian cells or under conditions of replication stress, and the assay can be performed in less than 2 weeks. Biotinylated Okazaki fragments are then captured on streptavidin beads and ligated to Illumina adapters before library preparation for Illumina sequencing. After size fractionation on a sucrose gradient, Okazaki fragments are concentrated and purified before click chemistry is used to tag the EdU label with a biotin conjugate that is cleavable under mild conditions. Briefly, cells are pulsed with 5-ethynyl-2′-deoxyuridine (EdU) to label newly synthesized DNA, and collected for DNA extraction. Here we describe a detailed protocol for isolating and sequencing Okazaki fragments from asynchronously growing mammalian cells, termed Okazaki fragment sequencing (Ok-seq), for the purpose of quantitatively determining replication initiation and termination frequencies around specific genomic loci by meta-analyses. coli is DNA Polymerase III (Pol III).The ability to monitor DNA replication fork directionality at the genome-wide scale is paramount for a greater understanding of how genetic and environmental perturbations can impact replication dynamics in human cells. The primary DNA polymerase for replication in E. Although only a few nucleotides are needed, the prokaryotic primers may be as long as 60 nt depending on the species.Īt least five prokaryotic DNA polymerases have been discovered to date. Therefore, a specialized RNA polymerase (RNAP’s do not have this limitation) known as primase is a part of the replisome, and reads creates a short RNA strand termed the primer for the DNA polymerase to add onto. DNA polymerases are unable to join two individual free nucleotides together to begin forming a nucleic acid they can only add onto a pre-existing strand of at least two nucleotides. However, before the DNA polymerases take positions, they need to be primed. Once oriC has been opened and the helicases have attached to the two sides of the replication fork, the replication machine, aka the replisome can begin to form. (C) Finally, the two temporarily broken ends of the DNA, which had been held closely in place by the enzyme. (B) This complete transection of the DNA (as opposed to the single-strand cut by type I topoisomerases) allows another part of the same DNA molecule to slip through the gap. (A) First, the enzyme binds to the DNA and initiates an endonuclease activity, cutting both strands of the DNA at that point. Detail of DNA topoisomerase type II action.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
